
Lifting the Lid on COVID
Lifting the Lid on COVID
To date, there are seven known coronaviruses that cause illness in humans, four of which only lead to mild to moderate disease.
Over the last two decades, three other coronaviruses have emerged from animal reservoirs and caused outbreaks across the globe, resulting in serious respiratory illness and in many cases death. Severe acute respiratory syndrome (SARS) emerged in 2002 as a result of the SARS coronavirus (SARS-CoV) and in 2012, Middle East respiratory syndrome coronavirus (MERS-CoV) was identified, and continues to cause sporadic and localized outbreaks.
The latest coronavirus strain to affect humans is severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), first reported in China in December 2019 and recognized as a global pandemic by the World Health Organization (WHO) in March 2020. This strain causes the disease termed COVID-2019, a respiratory illness that varies from mild flu-like symptoms to fatal lung disease.
How do Coronaviruses Infect Us?
For a virus to infect a living organism, it must invade cells to then multiply and continue infecting other hosts. The affected cells depend on the type of virus - coronaviruses specifically attack cells in the respiratory system. Following the SARS and MERS outbreaks and the more recent SARS-CoV-2, researchers have been dedicating their studies to finding out how exactly these viruses enter our cells, in the hope of finding a way to prevent it.
When coronaviruses enter the body, they must fuse with the host cell lipid membrane before delivering their genetic material. The fusion mechanism is not fully understood, but it is known that a so-called spike (S) protein - which is structurally categorized as a class I viral fusion protein, within the same group as influenza virus and HIV - facilitates binding and entry. Proteolytic cleavage of the S-protein exposes its fusion peptide (FP), which initiates membrane fusion. As a key component of how coronaviruses attack cells, locating and characterizing the FP is a priority for research groups, but its exact location has been disputed.
Important Analytical Tools
One technique researchers are using to help locate the coronavirus FP is electron paramagnetic resonance (EPR) spectroscopy. EPR is used to identify and quantify paramagnetic species (free radicals and transition metal ions) in solids, liquids, gases, cells, and in vivo - making it a popular analytical method for a range of applications, from nano-diamonds to cell membranes.
A study of the SARS-CoV FP used EPR (ELEXSYS™, Bruker BioSpin) for structural and functional analyses of peptide-lipid membrane interactions1. The study simulated the binding of different peptides isolated from SARS-CoV with synthetic lipid membranes, known as multilamellar vesicles (MLVs). The MLVs were labeled with a specific class of nitroxides called spin labels, which are sensitive to the local environment. The FP could be identified from the peptide mixture using membrane ordering as a parameter, which causes changes in the line width and shape of the EPR spectra. Using EPR, the group could identify and characterize the SARS-CoV FP.
SARS-CoV was chosen as a model due to the larger amount of functional data available, but the FP is highly conserved across the coronavirus family, so the findings of this study are applicable to all these viruses, including MERS-CoV and any novel viruses emerging from the animal reservoir, such as SARS-CoV-2 (COVID-19). Studies such as this continue to grow our understanding of how coronaviruses infect humans, and contribute to the end goal of preventing their spread.
For more information on our EPR spectroscopy solutions, please visit www.bruker.com/products/mr/epr.html
References:
- Lai AL, Millet JK, Daniel S, Freed JH and Whittaker GR (2017) The SARS-CoV Fusion Peptide Forms an Extended Bipartite Fusion Platform that Perturbs Membrane Order in a Calcium-Dependent Manner, J Mol Biol, 8;429(24):3875-3892.
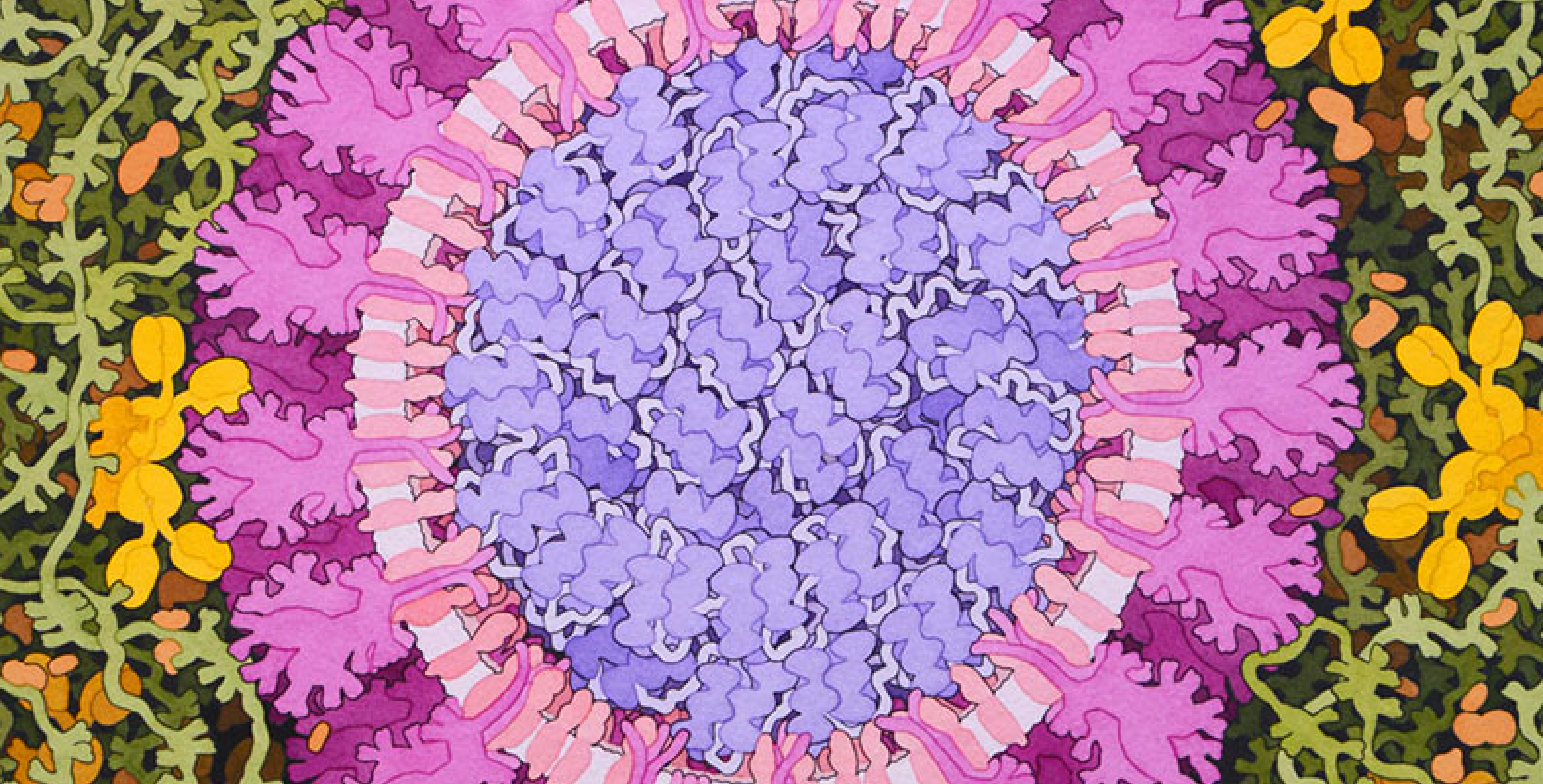
Acknowledgement: Illustration by David S. Goodsell, RCSB Protein Data Bank; doi: 10.2210/rcsb_pdb/goodsell-gallery-019
This painting depicts a coronavirus just entering the lungs, surrounded by mucus secreted by respiratory cells, secreted antibodies, and several small immune systems proteins. The virus is enclosed by a membrane that includes the S (spike) protein, which will mediate attachment and entry into cells, M (membrane) protein, which is involved in organization of the nucleoprotein inside, and E (envelope) protein, which is a membrane channel involved in budding of the virus and may be incorporated into the virion during that process. The nucleoprotein inside includes many copies of the N (nucleocapsid) protein bound to the genomic RNA.