Lipid Analysis Enabling Resistance Detection
Exploring lipidomics for rapid resistance detection
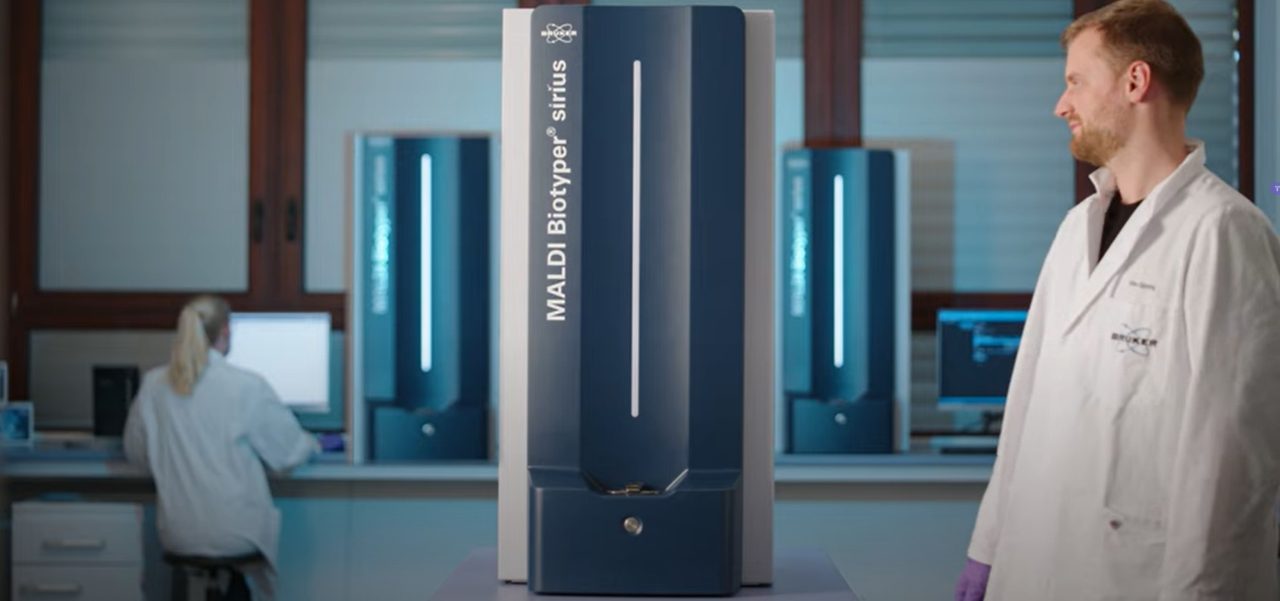
Analysis of microbial lipids enables fast resistance detection based on MALDI-TOF analysis of Lipid A modifications, on the MALDI Biotyper® sirius System, using negative ion mode.
Negative ion mode: unveiling new horizons in microbial analysis
Traditionally, lipid analysis with MALDI-TOF is performed in the negative ion mode, and now, this capability is seamlessly integrated into the MALDI Biotyper® sirius.
Unlock a powerful application in gram-negative bacteria — detecting Lipid A modifications and gaining insights into colistin resistance. The MALDI Biotyper® sirius, equipped with the MBT HT LipidART software module, makes this process straightforward and informative. Thanks to the user-friendly and efficient lipid extraction protocol provided by the MBT Lipid Xtract™ Kit, you can explore lipid analysis with ease.
The link between colistin resistance and Lipid A modifications
Colistin, one of our last lines of defence against gram-negative bacteria, is facing a growing threat due to bacterial resistance. This resistance often hinges on modifications to the bacterial lipopolysaccharides (LPS), with alterations in Lipid A playing a crucial role. These changes create a charge repulsion effect against colistin, rendering it ineffective in combating these resilient bacteria.
The modifications to Lipid A are diverse, including the addition of phosphoethanolamine (pEtN) and 4-amino-4-deoxy-L-arabinose (L-Ara4N) in various regions. These modifications lead to distinctive mass shifts in Lipid A.
Streamlined sample preparation
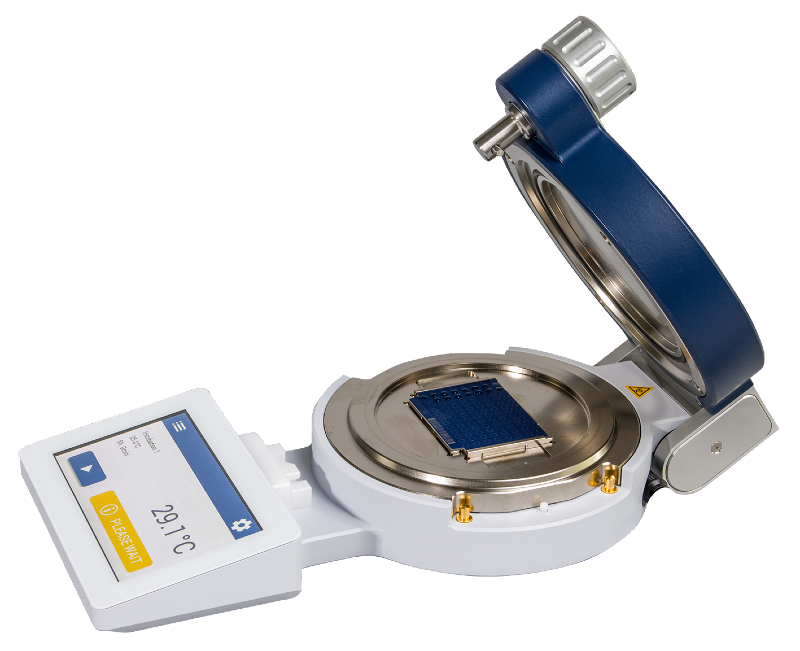
Sample preparation for lipid analysis can be done in just 15 minutes, with the MBT Lipid Xtract™ Kit. This efficient process involves lysing a small bacterial colony, followed by mixing with the Lipid Xtract matrix after a simple washing step. The resulting mixture can then be seamlessly transferred to an MBT Biotarget 96 for MALDI-TOF analysis in the negative ion mode, acquiring lipid spectra in the lower mass range.
Interpreting Lipid A spectra
Our dedicated software module simplifies the interpretation of the lipid MALDI-TOF spectra. With just a few clicks, you can extract valuable insights, distinguishing between chromosomal and plasmid-encoded resistance.
As the results are available after only 30 minutes starting from a colony, this assay enables testing many samples in a short time.
Join the ranks of visionary researchers who are revolutionizing microbial analysis
Lipid analysis for resistance detection requires:
A MALDI Biotyper® sirius System, operated in negative ion mode, and equipped with the MBT HT LipidART Module, for analysis of a sample prepared using the MBT Lipid Xtract™ Kit.
For Research use only. Not for use in clinical diagnostic procedures.
Not for sale in the USA. Please contact your local representative for availability in your country.