Application Note: The Application of NanoWizard ULTRA Speed AFM to Study Dynamics of Living KPG7 Fibroblasts
Dynamic measurement of live cells at near-physiological liquid conditions requires faster image acquisition than is typically achievable with conventional AFM imaging. Moreover, working with these often soft, topographically inhomogeneous materials is made more challenging by their weak and rapidly changing sample signals.
Nevertheless, imaging soft biological matter and rapidly changing, very active living cells is a fundamental part of studying dynamic cellular interactions. These events are typically impossible to observe with conventional AFM, but can be captured using the fast-scanning NanoWizard ULTRA Speed AFM due to its high spatiotemporal resolution, improved feedback response, and other leading-edge enhancements.
In this application note, we demonstrate these capabilities. The NanoWizard ULTRA Speed AFM is successfully used to monitor cytoskeletal reorganization and cell membrane protrusion events in living KPG7 fibroblasts in nearly physiological buffer conditions.
Readers can expect to learn about:
- AFM-based investigation of challenging biological/soft materials and systems;
- How fast-scanning AFM can be used and combined with advanced optical techniques to gain new information about cellular interactions that were previously impossible to observe; and
- The collection of dynamic measurements on living cells and direct observation of cell membrane dynamics with the NanoWizard ULTRA Speed AFM.
KEYWORDS: Dynamic Cellular Processes; Fast-Scanning AFM; Fluorescence Microscopy; High-Speed AFM; Fibroblasts; Live Cell Research
Introduction
The last three decades have seen the rise of the atomic force microscope (AFM) as an indispensable tool for high-resolution structural analysis of specimens ranging from single molecules [1] to complex biological systems [2]. Unlike other high-resolution imaging techniques, such as advanced electron microscopy and super-resolution optical microscopy, AFM remains the only tool that currently offers premium resolution of the analysed biological systems (proteins, cells, etc.) while being able to simultaneously acquire information about the sample’s mechanical properties at near physiological/native sample conditions. It also does not demand any sample modification and does not introduce preparation artefacts. Studying single macromolecule dynamics and the function of complex biological systems, such as individual living cells, requires a tool that can meet the requirements for both high spatial and temporal resolution [3]. Developments in the last 10-15 years have paved the way towards the application of ultra-small cantilevers, piezoactuator-based sample scanners, and optical beam deflection (OBD) detectors for studying of high-speed single molecule processes [4–6]. Such high-speed developments are hardly applicable for living cells due to the significantly reduced XY-scan (a few micrometres) and Z-scan size (less than a micrometre) [7]. The very recent fast-speed developments in tip-scanning AFMs make possible the successful structural analysis of a multitude of dynamic processes in cells such as exocytosis, vesicle transport, cytoskeleton reorganization, cell migration, each taking place on the timescale of seconds. Structurally resolving morphological surface changes and these cellular events is no longer fundamentally limited by the conventional optical diffraction limit, but rather in combination with the advantages of optical/fluorescence detection systems such as observed with correlative atomic force microscopy [8].
ULTRA Speed imaging of cells in liquid
The conventional AFM imaging of live cells in intermittent contact (and contact) mode is typically rather challenging due to the rather slow image acquisition times and relatively slow feedback being unable to cope with the rather soft and topographically inhomogeneous samples. The fast-scanning NanoWizard® ULTRA Speed AFM from JPK Instruments AG can now be applied for studying of weak and rapidly changing signals, such as dynamic cellular processes at near physiological liquid conditions.
This is made possible by the latest enhancements with one of the lowest noise scanners, very fast feedback algorithm and detection systems, which deliver the most accurate force control on delicate samples such as living cells and soft single molecules.
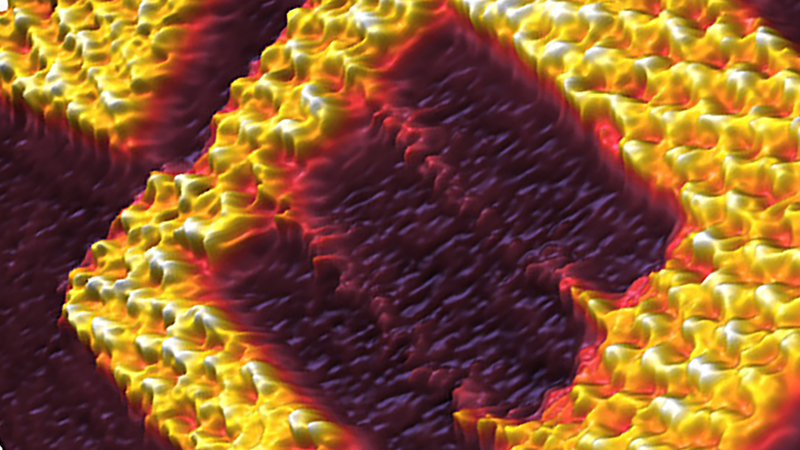
The example in Fig. 2 is given for a living KPG7 fibroblast, acquired in DMEM/F-12 cell culture medium with temperature control of 37 °C. It clearly resolves the filamentous structure of the underlying cytoskeleton. Ultra-Short Cantilever (USC from NanoWorld AG) with a resonant frequency of ~ 0.3 MHz and spring constant ~ 0.3 N/m were used for all of the measurements, carried out in a JPK PetriDishHeater™. The same imaging conditions were also applied to the rest of experimental examples.
Real-time cytoskeleton reorganisation
Successful dynamic measurements on living cells require a significant reduction in image acquisition times. In the example given below, the cell was imaged with a line-rate of 45 Hz and at resolution of 256x256 pixels, which results in ~ 5.7 s per frame. Figures 3 A-C show the phase channel of three consecutive frames that clearly show the cytoskeletal reorganisation taking place under the cell membrane of these quite active fibroblasts. The cell was imaged for about an hour at different speeds and scan sizes to verify that the tip-sample interaction is not leading to a specific cell retraction or introduction of further morphological artefacts.
In the case of rather complex macromolecular networks, it is often difficult to draw conclusions about the effective changes in the underlying cell cytoskeleton or of any associated migration events. By applying an external plugin for PIV it is possible to roughly visualise the displacements/force fields between the different images (Figure 3 D-E). This can be rather helpful with the visualisation of dynamic contractile processes arising from the filamentous actin reorganisation changes, as well as formation of focal adhesion points, as seen above.
Cell membrane dynamics
Another example of the application of fast AFM for studying biological systems with high temporal resolution is the direct observation of cell membrane dynamics. Living cells are constantly interacting with their surroundings, exchanging a number of molecules and signals. This is mostly associated with membrane turnover, e.g. membrane ruffling or vesiculation e.g. with internalisation of external molecules and vesicles (endocytosis), the release of metabolitic degradation products or the release of signalling molecules to other cells (exocytosis). The timescale of most of these processes can depend on the type of membrane fusion or secretion, and normally ranges for seconds to minutes [9], [10]. The example below shows a plausible exocytosis event on the timescale of 30-40 seconds, which is associated with a morphological change happening directly on the cell surface (Figure 4).
The budding vesicle can be clearly resolved in the phase images due to the much higher sensitivity of the phase channel to the rapidly changing and weak signals from the morphological differences in the cell surface. The same features can be clearly resolved in the height channel as well, which corroborates to the very good force control applied to study even such soft membrane structures. The height of the budding area gradually increased from 10 to about 25 nm.
LEARN MORE:
Conclusions
The NanoWizard® ULTRA AFM was successfully applied to monitor two examples of dynamic biological processes in living KPG7 fibroblasts in nearly physiological buffer conditions. Both cytoskeletal reorganization and cell membrane protrusion events were studied on the time scale of 0.1-0.2 frames per second (fps). Such events are typically impossible to observe with conventional AFM due to the rather low temporal resolution, as well as the slow feedback response being unable to resolve the rather heterogeneous structural features of rapidly changing and very active living cells. The imaging of soft biological matter now makes it possible to get beyond a single snapshot of cellular events, and gain additional information about the interaction between not only single cells, but also organelles and macromolecular protein complexes. The full compatibility of the system with high-resolution optical techniques for correlative microscopy opens new possibilities to combine its native high spatiotemporal resolution with the additional molecular information available from various molecular or immunostains. Future examples and studies will demonstrate the ability of the system to image dynamic biopolymerisation processes that underlie the formation of a number of key macromolecules taking place both in vivo and in vitro.
Acknowledgments
We would like to thank Roland Schwarzer and Andreas Herrmann from the group of Molecular Biophysics at the Humboldt University Berlin for providing the KPG7 fibroblasts for experiments.
References
- D. J. Muller, “AFM: a nanotool in membrane biology.,” Biochemistry, vol. 47, no. 31, pp. 7986–7998, Aug. 2008.
- A. Berquand, C. Roduit, S. Kasas, A. Holloschi, L. Ponce, and M. Hafner, “Atomic Force Microscopy Imaging of Living Cells,” Microscopy Today, p. 8, 2010.
- R. Tomer, K. Khairy, F. Amat, and P. J. Keller, “Quantitative high-speed imaging of entire developing embryos with simultaneous multiview light-sheet microscopy.,” Nat Methods, vol. 9, no. 7, pp. 755–763, Jul. 2012.
- T. Ando, N. Kodera, E. Takai, D. Maruyama, K. Saito, and A. Toda, “A high-speed atomic force microscope for studying biological macromolecules.,” Proc Natl Acad Sci U S A, vol. 98, no. 22, pp. 12468–12472, Oct. 2001.
- L. Bozec A. Ulcinas D. J. Engledew M. Antognozzi M. A. Horton L. M. Picco and M. J. Miles, “Breaking the speed limit with atomic force microscopy,” Nanotechnology, vol. 18, 2007.
- M. B. Viani, L. I. Pietrasanta, J. B. Thompson, A. Chand, I. C. Gebeshuber, J. H. Kindt, M. Richter, H. G. Hansma, and P. K. Hansma, “Probing protein-protein interactions in real time.,” Nat Struct Biol, vol. 7, no. 8, pp. 644–647, Aug. 2000.
- T. Ando, “High-speed atomic force microscopy coming of age,” Nanotechnology, vol. 23, no. 6, p. 062001, 2012.
- A. Monserrate, S. Casado, and C. Flors, “Correlative Atomic Force Microscopy and Localization-Based Super-Resolution Microscopy: Revealing Labelling and Image Reconstruction Artefacts.,” Chemphyschem, Nov. 2013.
- J. Klingauf, E. T. Kavalali, and R. W. Tsien, “Kinetics and regulation of fast endocytosis at hippocampal synapses.,” Nature, vol. 394, no. 6693, pp. 581–585, Aug. 1998.
- S. W. Schneider, K. C. Sritharan, J. P. Geibel, H. Oberleithner, and B. P. Jena, “Surface dynamics in living acinar cells imaged by atomic force microscopy: identification of plasma membrane structures involved in exocytosis.,” Proc Natl Acad Sci U S A, vol. 94, no. 1, pp. 316–321, Jan. 1997.
SEE THE LATEST GENERATION NANOWIZARD: