Beta-Lactamase Activity Detection
Revealing beta-lactamase activity
To aid the fight against resistant bacteria, Bruker has developed the MBT STAR®-BL assays. Extending the capabilities of the MALDI Biotyper® beyond the identification of microorganisms, these assays provide rapid assessment of beta-lactamase activity in bacterial isolates. Dedicated MBT STAR® reagent kits and software modules facilitate straight-forward evaluation of results.
Detecting beta-lactamase activity with the MALDI Biotyper®
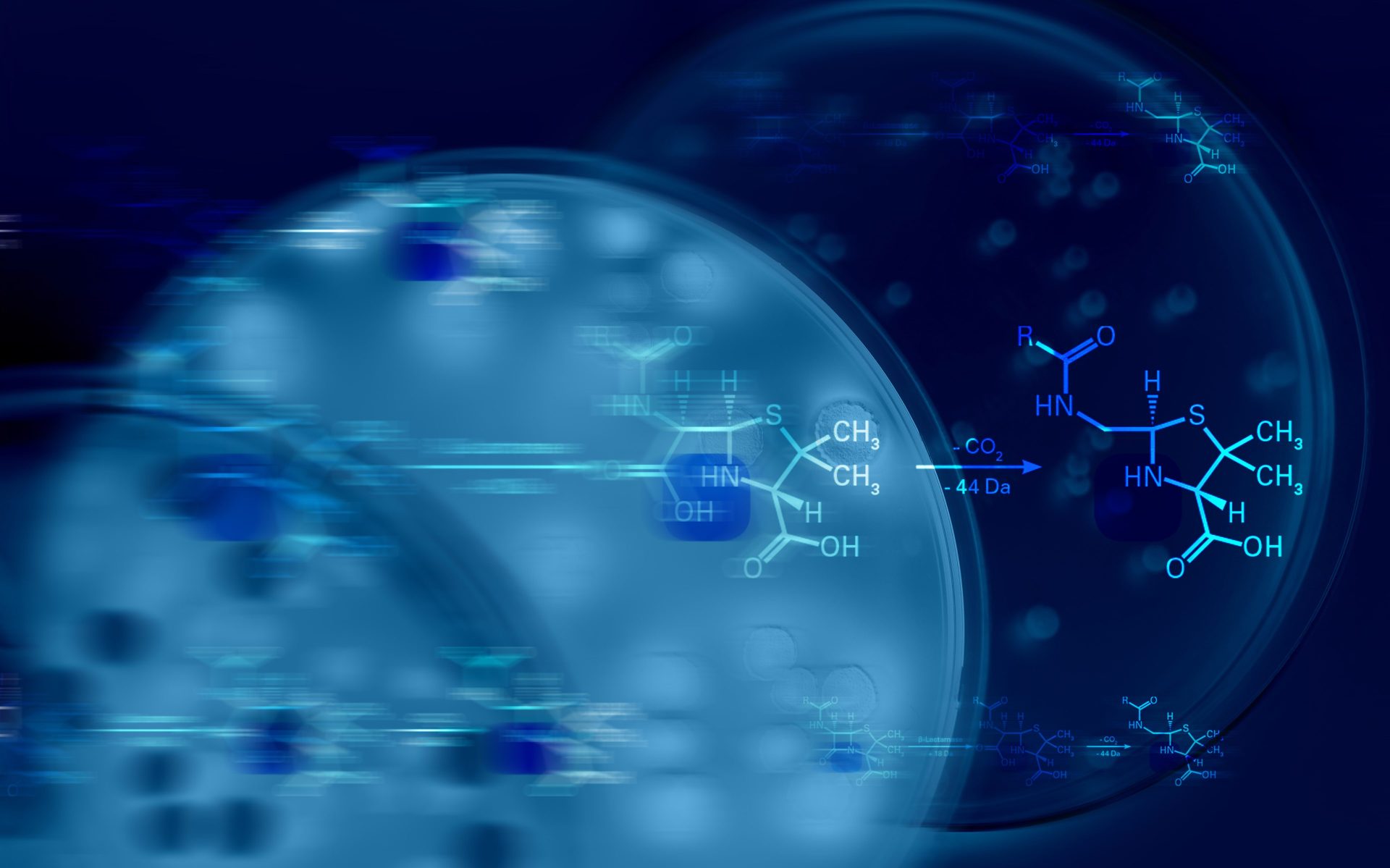
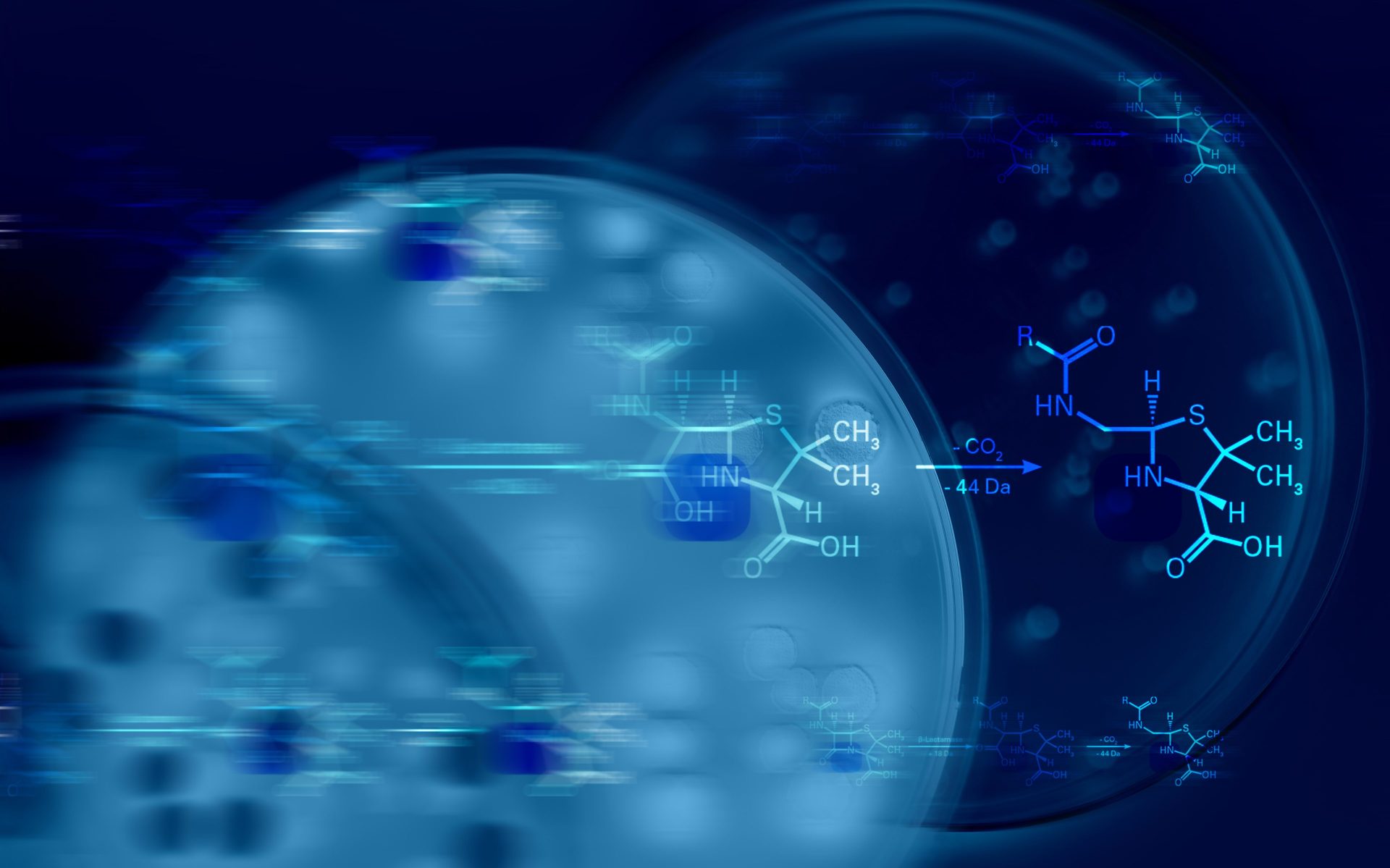
To analyze beta-lactamase activity, bacterial samples are co-incubated with a beta-lactam antibiotic. Bacteria possessing beta-lactamase activity can effectively break down the antibiotic molecule. This enzymatic action results in a detectable mass shift of the molecule. That mass shift, being the core of our methodology, is precisely measured on the MALDI Biotyper®.
Rapid results, easy execution
Following incubation, assay samples are deposited onto a MALDI target plate, and spectra of the antibiotic are swiftly acquired on the MALDI Biotyper®. The dedicated MBT HT STAR®-BL software module automatically computes the combined signal intensities of hydrolyzed and intact antibiotic molecules, presenting the data as a comprehensible plot for instant evaluation.
Our assays are designed for your convenience, delivering fast results and requiring only 30-60 min of incubation. Utilize the user-friendly MBT STAR®-Carba and MBT STAR®-Cepha Kits to swiftly detect carbapenemase and cephalosporinase activity, respectively. Whether you're starting with a plate culture or a Sepsityper® pellet, our solutions empower you to stay ahead in the fight against antibiotic resistance.
Dedicated kits and software module for detection of carbapenemase and cephalosporinase activity
The MBT STAR®-Carba assay is designed for the swift detection of prevalent Class A, B, or D carbapenemase activity in gram-negative Enterobacteriaceae, Pseudomonas spp., and Acinetobacter spp. This assay equips microbiologists with the means to rapidly identify bacteria and assess their carbapenemase activity seamlessly on the MALDI Biotyper® - all on a single platform.
Employ the MBT STAR®-Cepha assay in the fight against cephalosporin resistance. Swiftly identify resistances to 3rd generation cephalosporins after a positive blood culture alert. The MBT STAR®-Cepha assay identifies the characteristic enzymatic hydrolysis of the beta-lactam ring in 3rd generation cephalosporin antibiotics.
The MBT STAR®-Cepha assay enables detecting the most actively expressed extended-spectrum beta-lactamases (ESBL) and AmpC beta-lactamases in multi-drug-resistant (MDR) gram-negative Enterobacterales:
ESBL:
- e.g. plasmidic TEM, SHV, and CTX-M
AmpC:
Chromosomal and plasmidic
Inducible or overexpressed resistance genes
e.g. AmpC, FOX, LAT, DHA, and CMY
Users receive a comprehensive report, featuring assay results and clear, color-coded indicators, simplifying the interpretation of enzyme activity status.
Selective Testing of Antibiotic Resistance (STAR) solutions fitting to your microbiology lab
Bruker’s MBT STAR® solutions are available as IVD-CE workflows, or for Research Use Only. Please contact your local representative for availability in your country.
MBT HT STAR®-BL IVD Module
This module is for evaluation of beta-lactamase activity detection assays using the MBT STAR®-Carba IVD Kit and/or MBT STAR®-Cepha IVD Kit.
For professional use only. Not for sale in the USA.
MBT HT STAR®-BL Module
This module is for evaluation of beta-lactamase activity detection assays using the MBT STAR®-Carba Kit and/or MBT STAR®-Cepha Kit.
For Research use only. Not for use in clinical diagnostic procedures.