Imaging Extracellular Vesicles with Super-Resolution Microscopy
TABLE OF CONTENTS:
Single-molecule localization microscopy (SMLM) enables high-resolution imaging of extracellular vesicles (EVs). It allows highly accurate quantification of their size, molecular content, and cellular uptake/cargo transfer processes.
In doing so, SMLM:
- Overcomes the limitations of traditional EV imaging methods, and
- Achieves the accuracy and specificity researchers need to expand our understanding of EVs, their role in intercellular communication, and their contribution to pathologies.
Continue reading, contact us, or see the Vutara VXL SMLM microscope to learn more.
The Benefits of Single Molecule Localization Microscopy for EV Analysis
EVs are small, lipid bound objects excreted by cells. With sizes in the low hundreds of nanometers, very high resolution microscopy techniques are needed to directly image them.
Single molecule localization microscopy (SMLM) techniques like dSTORM or DNA-PAINT provide roughly 20 nm resolution radially and 50 nm axially. This resolution provides the ability to determine if a component is EV cargo, external to the EV, or inserted into the membrane. SMLM can show if labeled cargoes or components are distributed uniformly or asymmetrically. With dSTORM or DNA-PAINT imaging we can co-label for multiple targets of interest and correlate their presence on the same EV.
3D volumetric imaging of EVs
Even when purified out of biomaterials and affixed to a coverslip, EVs have three-dimensional structure. A cross section through even a roughly spherical object may not give you the full story. With our ability to image intrinsically in 3d due to our patented biplane technology, you can characterize your EVs in all three spatial dimensions. 3D dSTORM imaging allows for accurate measurements of volume, label distribution, and volumetric co-localization of analysis, more accurately reflecting the physical reality of your sample.
Multi-color imaging of EVs
The Bruker Vutara VXL supports multi-color imaging of extracellular vesiclesWith our powerful lasers, dSTORM dyes blink quickly, improving detection sensitivity and reconstruction quality, by eliminating wasteful pre-bleaching steps.
Because we fit a measured biplane point spread function to your data, when we measure the point spread function as part of our calibration, we also correct for the chromatic aberration of the optical system. If any residual chromatic aberration remains, it can be corrected for in software. If these methods are deemed inadequate for objects of EV size, we also offer a dual camera spectral demixing system, which allows for the simultaneous localization of spectrally similar dyes (such as CF647 and CF680), which prevents chromatic aberration effects by minimizing the difference in diffraction. This spectral demixing system could also be used to detect an additional label.
If dSTORM doesn’t offer enough labels, sequential labeling DNA-PAINT and/or antibody stripping and re-staining approaches are possible without removing the sample from the instrument with our PlexFlo fluidics units.
Study EVs in biological context
EVs are signal carriers. Purified analysis of EVs may advance our understanding of their composition, population and structure, but doesn’t address their targets, or interactions with cells tissues or organs. Because of the Bruker Vutara VXL’s Biplane, it can perform 3D Single Molecule Localization microscopy at depth.
If you want to understand how your EVs or synthetic lipid nanoparticles interact with your model organism’s physiology, the Vutara VXL can be used to directly image this. Simply label your EVs, expose your model organism to them, then fix, section and stain for your cell type, receptor or structure of interest, and image them interacting with tissue
The Bruker Vutara VXL can image up to 50 microns into tissue, organoids, organs or whole model organisms, allowing the study of EVs in vivo at sub-extracellular vesicle super resolution.
Study EV biogenesis
Extracellular vesicles are secreted by and interact with cells. While fixed, stained experiments can answer many biological questions, directly imaging the dynamics of EV secretion, motion and uptake could open new, promising avenues for research.
The Bruker Vutara VXL is an ideal platform for live cell super resolution microscopy, supporting both single particle tracking and PALM.
Automated EV analysis
Purified EVs can be stained in solution and affixed to coverslips for batch and bulk analysis. Imaging high numbers of EVs provides power for statistical analysis. Because purified EV analysis is a routine process, we have implemented an automated analysis workflow. Here are the steps:
- Collect many fields of view - Select one or more fields of view to image. Each field of view should contain hundreds to thousands of EVs.
- SRX software localizes your data in real time, on your PC - SRX runs all analysis steps on your PC. Localization is performed locally and in real time. No worries about internet connection and transfer of potentially protected or proprietary information over the internet required.
- Merge your fields of view - Review your data and merge the fields of view that correspond to your experimental group. If additional data is needed, you can collect more.
- Run automated EV analysis - Run our automated EV analysis workflow with a single button push.
- Review the results - Your EV report lists relevant info about your experiment: For each positivity class size distribution and count information is displayed.
- Plot and adjust parameters - Plot, filter, review and export parameters for your EVs by positivity class, or filter your particles to improve on our robust default analysis parameters.
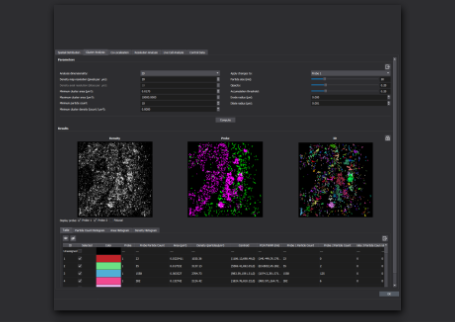
Our automated EV analysis workflow generates a report detailing the results of your experiment:
Our New, automated EV analysis workflow automatically generates a report detailing the results of your experiment.
From top of image:
- The fields of view you selected for analysis, along with the number of EVs detected in each field of view
- All detected positivity classes
- The total number of EVs detected in each class
- Size distribution for each positivity class
- The distribution of size for all detected EVs in your experiment
Answer Key Questions in EV Research
Resolving the size and composition of single EVs is critical to answering open questions in the study of EVs and understanding their potential therapeutic and clinical uses.
Super-resolution microscopy can distinguish between membrane proteins and internal cargo. It can determine the composition of EV cargo. It uniquely supports contemporary EV research focused on: (1) exploring the role EVs play in cellular communication and (2) the mechanisms and molecules behind EV deployment, release, uptake, and impact on the biochemical function and behavior of cells.
Researchers exploring these topics primarily seek to answer two questions:
- What type of EV is a particular cell making and releasing? and
- Where do those EVs go (e.g., what types of recipient cell(s) do they enter and what are the consequences of their uptake into those cell types)?
To a large extent, EV subgroups are determined by particle size, which may also play a role in cellular uptake. For example, the upper size limit for clathrin-mediated endocytosis of EVs is near 200 nm. Accurate and reproducible nanometer-scale measurements across all EV subgroups are essential to characterizing EVs and advancing our understanding of EV biology.
SMLM offers a uniquely reliable and practical method for sizing EVs. It enables researchers to easily differentiate and classify EV particles and calculate particle size distribution (PSD).
SMLM is a unique EV imaging method capable of:
- Sizing EVs down to 20 nm (or slightly less) — a capability critical to the characterization of exosomes;
- Accurate differentiation and measurements across a wide range of particle sizes; and
- Imaging and analyzing vesicles in isolation or in cells (in culture or tissue).
Molecular cargo content is a key point of interest in leading EV research; the effect EV uptake has on the biochemical function and behavior of destination cells is directly related to the composition of its molecular cargo. As a result, highly accurate molecular content/composition characterization is essential for exploring questions about the mechanisms of EV-mediated cellular communication.
SMLM provides highly accurate content data for single EVs
Using immunofluorescent labeling to target membrane proteins and cargo, SMLM can both visualize and quantify specific membrane proteins and determine the composition of a given vesicle’s molecular cargo (e.g., proteins, DNA, RNA, and lipids) — information that traditional methods of EV imaging are unable to determine on a single EV level.
Given that EVs can contain various cargos, ongoing research will involve methodologies such as proteomics or sequencing used in conjunction with super-resolution microscopy. These combined protocols will be especially relevant to discovering the complete composition of EVs generated by specific cells.
The mechanisms contributing to EV uptake and the biology underlying cargo transfer between cells may have significant implications for disease pathologies. Our understanding of these mechanisms has, until now, been constrained by the capabilities of traditional EV measurement techniques.
SMLM provides the consistency, resolution, and single-molecule sensitivity needed to detect, track, and study EV uptake and cargo transfer.
Its capabilities include:
- Monitoring the uptake of EVs of known composition into cells;
- Tracking EV release and uptake from cells of interest to target cells;
- Identifying EV components that promote their uptake and functional cargo transfer; and
- Visualizing and analyzing the uptake and processing mechanisms of specific EVs and EV subpopulations, differentiated by size and molecular cargo content.
These activities can provide insight into the mechanisms and molecules that initiate and impact selectivity for entry, entry processes, and the fate of EVs once they enter cells. Such insights may contribute to the development of EV-based therapeutics for various pathologies (e.g., cancer metastasis, viral infections, and immune responses) with greater effectiveness than current therapeutic options.
EV count and cargo might provide important information about the pathophysiology of various diseases. As a result, EVs have the potential to serve as excellent biomarkers, enabling researchers to:
- Clarify the underlying pathogenic mechanisms of diseases;
- Screen and diagnose diseases, particularly those in the early stages of pathogenesis and cancers of unknown origin; and
- Develop new and better therapeutic options.
SMLM may be helpful in revealing actionable biomarkers
With its unique combination of capabilities — particularly to detect, classify, size, count, track, and determine the contents of EVs at the single-vesicle level — SMLM is an ideal EV research method for the discovery of EV-based biomarkers.
Frequently Asked Questions
Core EV Characterization Questions
The Vutara VXL can size EVs down to 20 nm or less, which is a critical range for characterizing exosomes that traditional light-scattering techniques often struggle to resolve accurately.
Unlike light-scattering methods (such as NTA) that may confuse EVs with lipoproteins or large unilamellar vesicles (LUVs), the Vutara VXL uses fluorescence-based SMLM to specifically identify and distinguish specifically labeled EVs from contaminants.
The Vutara VXL can assess the cargo content (proteins, RNA, lipids) of single vesicles, both in aggregate in purified samples, and in interaction with target cells or in tissue section. The system’s single-molecule sensitivity allows for the characterization of composition and how it affects the biochemical behavior of recipient cells.
Questions on Biological Dynamics
The Vutara VXL can be used to monitor the release or uptake of labeled EVs of known composition from or into target cells, visualizing the internal processing these signal carriers.
The Vutara VXL can image EVs in tissues. If labeled EVs are applied to a research organism, those EVs can be subsequently imaged in fixed, stained tissue sections of interest, allowing study of EV interactions with organs.
Technical Implementation for EV Research
Yes, by utilizing the PlexFlo microfluidics unit, researchers can perform essentially unlimited multiplexed imaging to identify multiple biomarkers (e.g., CD81, CD63, CD9) on the same individual vesicles.
The system's bi-plane technology allows for 3D imaging up to 100 µm deep, enabling the study of EVs in their native environment within tissue slices, organoids, or model organisms.
You can read more about the Vutara VXL system, download the brochure, or contact a Bruker expert to discuss your specific measurement requirements.