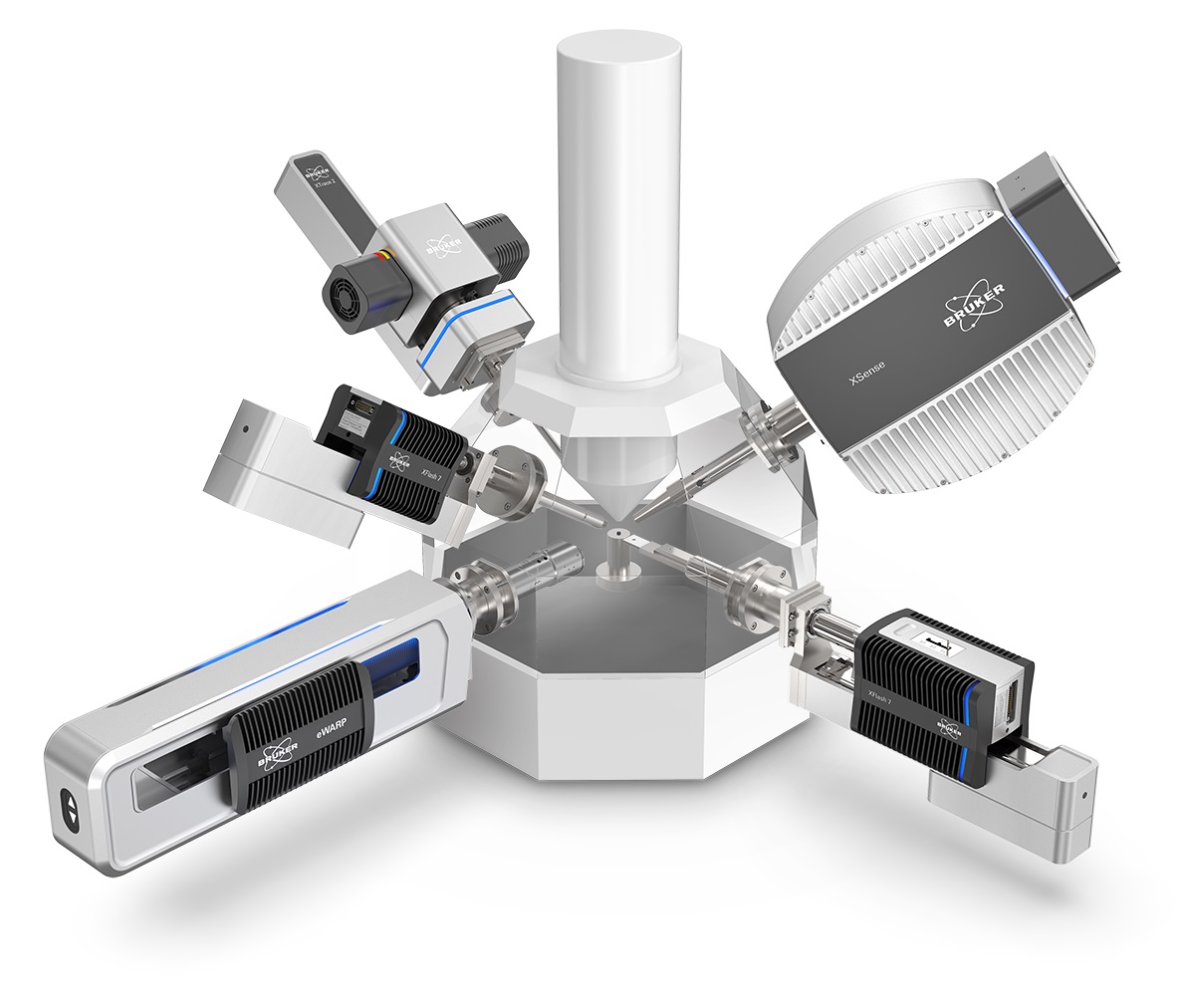
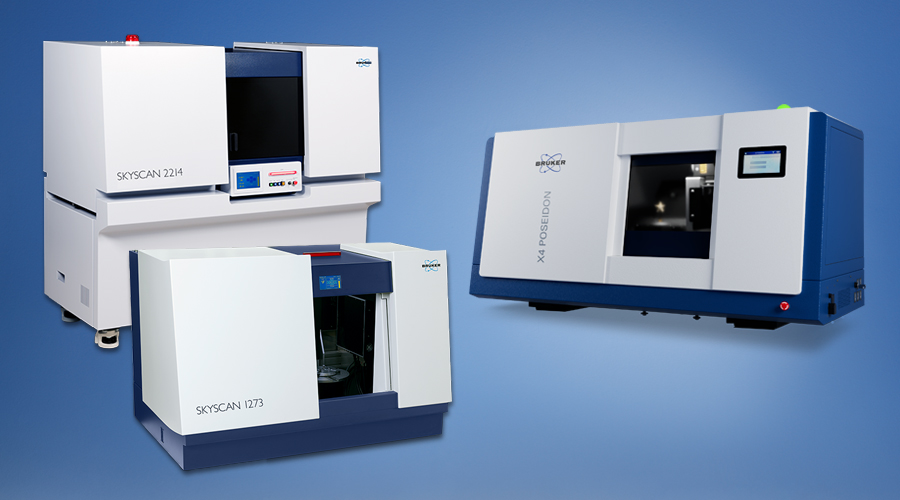
Microscopes
Enabling discovery at the molecular, cellular, microscopic and nanoscopic levels
Advancing Discovery with High-Performance Microscopy Solutions
Bruker's world-leading microscopy systems are helping scientists and engineers to make breakthrough discoveries and develop new applications and products that improve nearly all aspects of our world. Our microscopes enable scientists to explore life and materials at the molecular, cellular and microscopic levels. In close cooperation with our customers, these systems are driving innovation, improved productivity, and customer success in life science and materials research.